Recent discoveries reveal that soil microbes possess a remarkable form of ecological memory—communities repeatedly exposed to drought cycles recover faster and support plant growth more effectively than their unexposed counterparts. This breakthrough transforms how ecologists approach Soil Microbe Memory in Drought-Resilient Biodiversity Surveys: Protocols for Ecologists Facing 2026 Climate Extremes, offering a powerful new lens for quantifying ecosystem resilience through environmental DNA (eDNA) analysis and microbial community profiling. As climate extremes intensify across 2026, understanding and leveraging this microbial memory becomes essential for accurate biodiversity net gain assessments and long-term habitat restoration success.
Key Takeaways
- Soil microbes develop ecological memory from repeated drought exposure, enabling faster recovery and enhanced plant support during future stress events 🌱
- Plant diversity directly strengthens microbial resilience, with higher species richness (1-60 species gradients) improving microbial stress tolerance through increased carbon availability
- eDNA protocols can quantify microbial memory as a measurable resilience metric for biodiversity baselines, particularly valuable for biodiversity net gain planning
- Root depth architecture influences microbial performance, with deep-rooted perennials (reaching 2.5m) promoting greater microbial biomass and carbon cycling at depth
- Synthetic microbial communities (SynComs) show promise for accelerating post-drought recovery, with 15-species assemblages improving all plant growth traits during rewatering phases
Understanding Microbial Ecological Memory in Drought-Adapted Ecosystems
What Is Soil Microbe Memory?
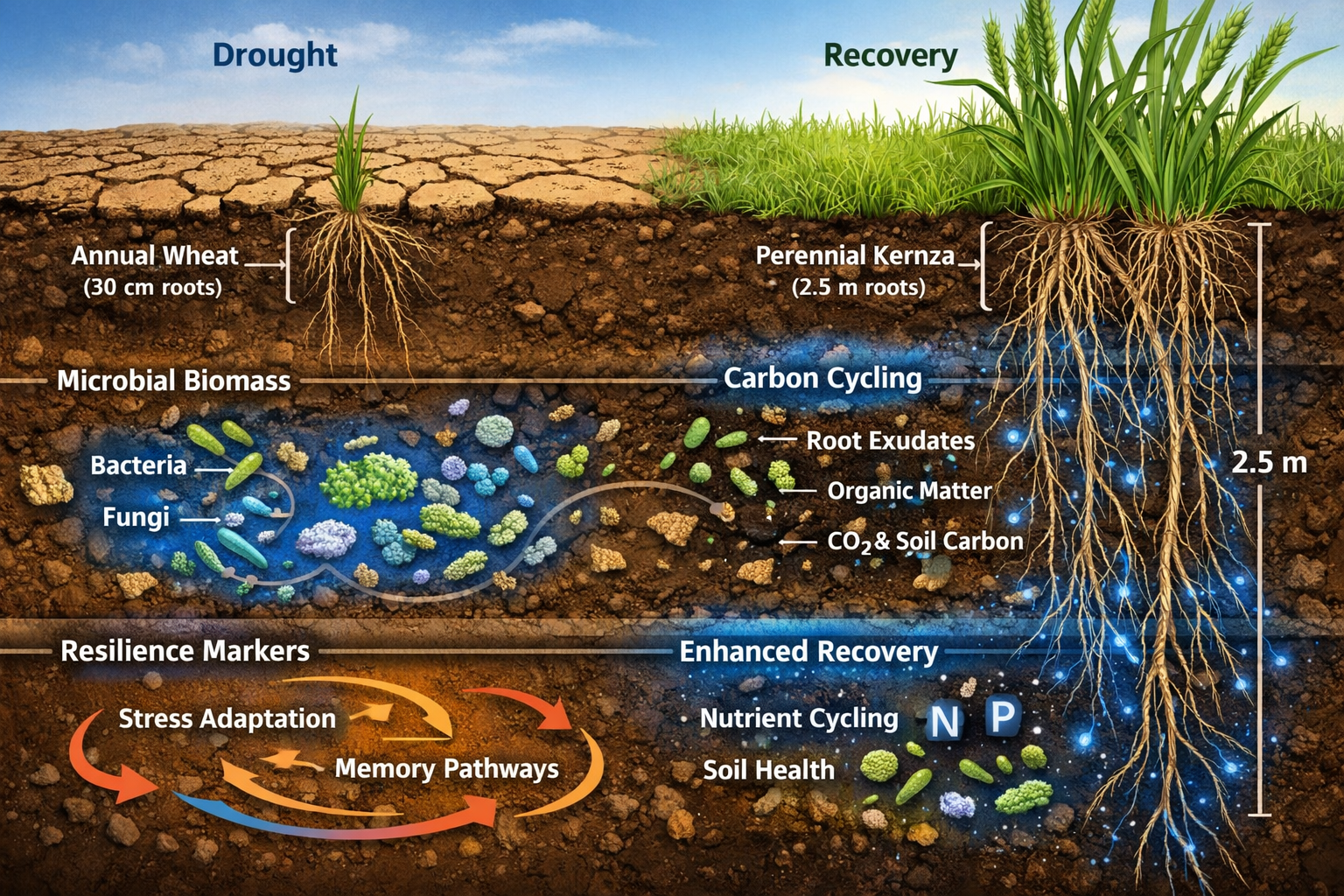
Soil microbial communities exhibit a phenomenon scientists call ecological memory—the ability to retain information about past environmental conditions and respond more effectively to similar future stresses. Research presented at the 2026 European Geosciences Union conference demonstrates that soil communities frequently exposed to drying-rewetting cycles recover growth more rapidly and exhibit sharper respiration peaks compared to less-adapted communities, which show delayed recovery responses.[1]
This memory manifests through several mechanisms:
- Compositional shifts: Drought-tolerant microbial species become dominant after repeated stress exposure
- Metabolic priming: Enzymatic pathways remain partially activated, enabling faster response times
- Structural adaptations: Formation of protective biofilms and spore-producing populations
- Genetic selection: Enrichment of stress-resistance genes within the community
The practical implications for ecologists conducting biodiversity impact assessments are profound. Sites with climate-adapted microbial communities possess inherently greater resilience—a factor that should influence baseline valuations and restoration target-setting.
The Plant-Microbe Resilience Connection
Higher plant species richness positively affects both microbial resistance (ability to withstand stress) and resilience (speed of recovery). Studies testing gradients from 1 to 60 plant species found that diverse plant communities increase high-quality carbon availability in soils, which directly supports microbial stress tolerance.[1]
This relationship creates a positive feedback loop:
- Diverse plant communities produce varied root exudates and litter chemistry
- Microbial communities develop functional diversity to process these resources
- Functionally diverse microbes better withstand environmental fluctuations
- Resilient microbes support plant recovery during and after drought
- Recovered plants maintain diverse microbial populations
For ecologists designing biodiversity net gain strategies, this means plant diversity targets must consider their underground microbial partners. A monoculture restoration, even of native species, will fail to develop the microbial resilience necessary for long-term climate adaptation.
Root Architecture and Microbial Performance at Depth
The vertical dimension of plant-microbe interactions proves equally critical. Research comparing shallow-rooted annuals (wheat) with deep-rooted perennials (kernza, reaching 2.5m depth) reveals that longer root systems promote greater microbial biomass and enhanced carbon cycling at depth.[1]
| Root System Type | Maximum Depth | Microbial Biomass Impact | Carbon Cycling Benefit |
|---|---|---|---|
| Annual wheat | 30-60 cm | Moderate (surface-focused) | Limited to topsoil |
| Perennial grasses | 1-1.5 m | High (distributed) | Extended through profile |
| Deep perennials (kernza) | 2-2.5 m | Very high (deep zones) | Significant at depth |
This depth effect matters because deeper soil layers provide refugia during surface drought, allowing microbial populations to persist and recolonize upper layers during recovery. Ecologists should therefore prioritize deep-rooted species in restoration plans, particularly for sites facing projected increases in drought frequency.
Protocols for Integrating Soil Microbe Memory into Biodiversity Surveys

eDNA Sampling Protocols for Microbial Community Assessment
Environmental DNA (eDNA) analysis provides a powerful, non-invasive method for characterizing soil microbial communities and their drought-memory signatures. The following protocol outlines best practices for Soil Microbe Memory in Drought-Resilient Biodiversity Surveys: Protocols for Ecologists Facing 2026 Climate Extremes:
Field Sampling Protocol
Timing considerations:
- Sample during both drought and recovery phases to capture memory responses
- Coordinate with seasonal moisture patterns (spring recovery, summer stress, autumn stabilization)
- Repeat sampling across multiple years to document memory development
Spatial sampling design:
- Establish 10m × 10m survey plots within each habitat type
- Collect 5-10 composite samples per plot (combining 3-5 subsamples each)
- Sample at multiple depths: 0-10cm, 10-30cm, 30-60cm, and >60cm where possible
- Include reference samples from undisturbed analog sites with known drought history
Sample collection technique:
- Use sterile soil corers (2.5cm diameter minimum)
- Collect 50-100g per composite sample
- Store immediately on ice or in portable -20°C freezer
- Process within 24 hours or freeze at -80°C for later analysis
Laboratory Analysis
Modern eDNA analysis for microbial communities involves:
- DNA extraction: Use commercial kits optimized for soil (e.g., PowerSoil, DNeasy)
- Amplicon sequencing: Target 16S rRNA (bacteria/archaea) and ITS (fungi) regions
- Shotgun metagenomics: For functional gene profiling (stress-response genes, antibiotic resistance markers)
- Bioinformatic analysis: Quantify diversity metrics, functional potential, and drought-memory indicators
Quantifying Microbial Memory as a Resilience Metric
To incorporate microbial memory into biodiversity net gain calculations, ecologists need standardized resilience metrics. The following framework adapts traditional biodiversity scoring to include microbial dimensions:
Microbial Resilience Index (MRI) Components:
📊 Community Composition Score (0-3 points)
- Proportion of drought-tolerant taxa (>50% = 3 points, 30-50% = 2 points, <30% = 1 point)
- Presence of keystone resilience species (identified through literature and regional reference data)
📊 Functional Capacity Score (0-3 points)
- Abundance of stress-response genes (heat shock proteins, osmolyte synthesis pathways)
- Diversity of carbon-cycling enzymes (cellulases, laccases, peroxidases)
📊 Recovery Potential Score (0-4 points)
- Ratio of fast-growing to slow-growing taxa (r-strategists vs. K-strategists)
- Spore-forming bacterial abundance
- Mycorrhizal fungal colonization potential
Total MRI = 0-10 points, which can be integrated into broader habitat condition assessments and biodiversity unit calculations.
Integrating Microbial Data with Traditional Biodiversity Surveys
Microbial memory assessments should complement, not replace, traditional above-ground biodiversity surveys. The integration process involves:
Baseline Assessment Phase:
- Conduct standard habitat condition assessments (vegetation structure, species composition)
- Simultaneously collect soil samples for eDNA analysis
- Document site drought history (meteorological records, vegetation stress indicators)
- Calculate preliminary biodiversity units using conventional methods
Enhanced Resilience Scoring:
- Apply MRI scores as a condition modifier (±10-20% adjustment to habitat distinctiveness)
- Weight adjustments based on projected climate scenarios for the site
- Document microbial memory as a "future-proofing" factor in biodiversity net gain reports
Monitoring and Adaptive Management:
- Establish 5-year monitoring protocols with annual eDNA sampling
- Track microbial community convergence toward target reference conditions
- Adjust management interventions based on microbial recovery trajectories
This integrated approach ensures that biodiversity plans account for both visible and invisible components of ecosystem resilience.
Leveraging Synthetic Microbial Communities for Enhanced Recovery
The SynCom Approach to Drought Resilience
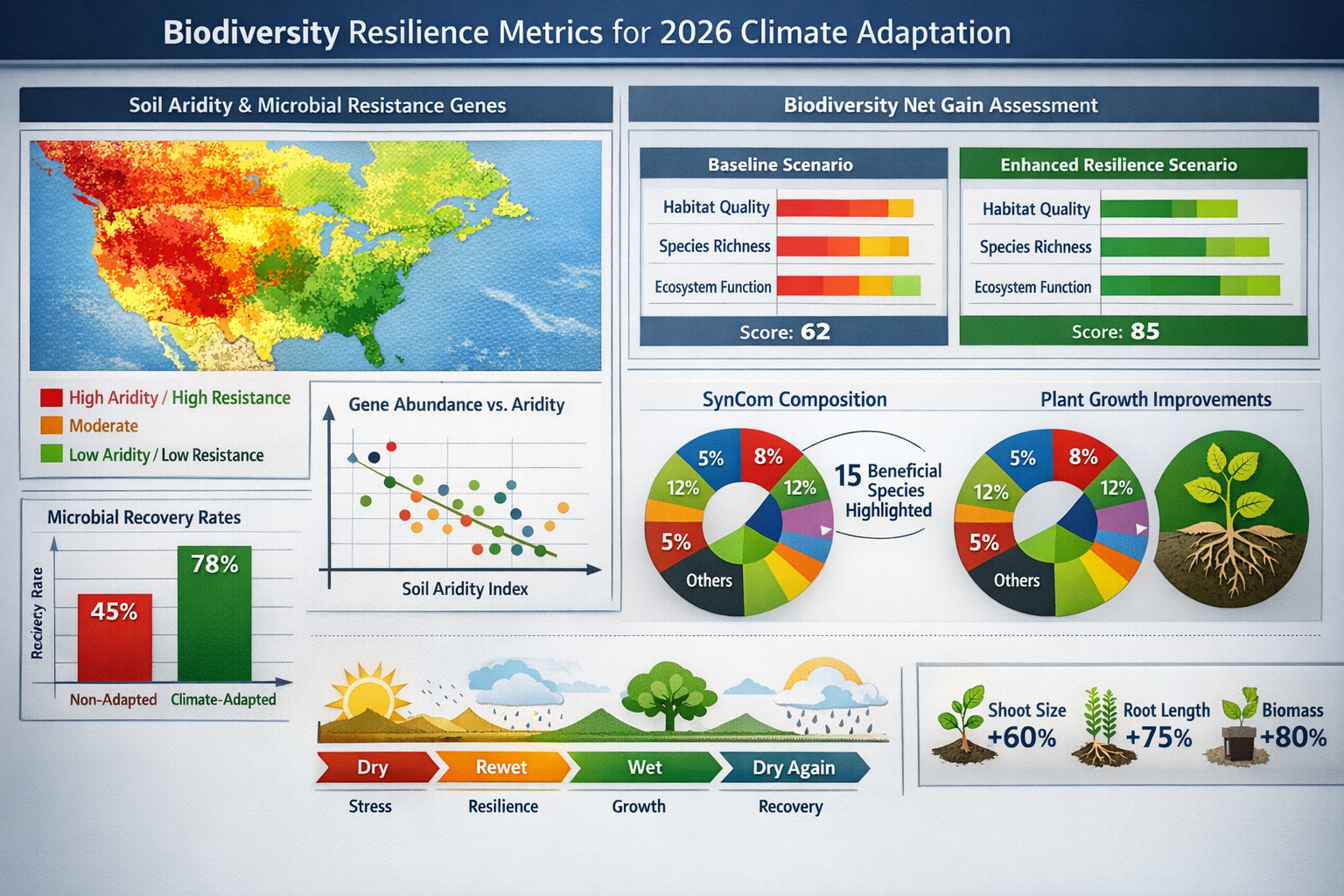
Recent research on Brachypodium distachyon grass demonstrates that synthetic microbial communities (SynComs) composed of 15 carefully selected rhizosphere microbe species significantly improve all plant growth traits during drought-recovery conditions.[2] After three weeks of recovery, SynCom-treated plants showed improvements in:
- Shoot size and biomass 🌿
- Root length and architecture 🌱
- Overall plant weight ⚖️
- Leaf count and area 🍃
Critically, these microbe treatments provided greater benefits during the rewatering phase than during initial drought exposure, indicating their primary value lies in facilitating ecosystem recovery from extreme events rather than preventing stress damage.[2]
Microbial Recruitment Signals Under Stress
Plants actively recruit beneficial microbes through root exudates—chemical signals released into the rhizosphere. Under drought and salt stress, specific microbes accumulate near the root tip (the actively growing meristematic region), suggesting plants send targeted chemical signals to attract growth-supporting microbes precisely where they're needed most.[2]
This selective colonization has important implications for restoration ecology:
- Inoculation timing matters: Apply SynComs during early establishment phases when root systems are actively developing
- Stress-adapted inocula outperform generic products: Select microbial strains with demonstrated drought-memory characteristics
- Root architecture influences recruitment success: Deep-rooted species provide more colonization opportunities throughout the soil profile
Practical Applications for Habitat Creation
For ecologists implementing biodiversity net gain projects, SynCom applications offer several advantages:
On-site habitat creation:
- Accelerate establishment of drought-resilient plant communities
- Reduce irrigation requirements during vulnerable establishment phases
- Improve survival rates of translocated vegetation
- Enhance soil carbon sequestration through promoted microbial activity
Off-site habitat banking:
- Develop climate-adapted habitats more rapidly, reducing time-to-target-condition
- Increase reliability of habitat banking investments
- Provide measurable resilience upgrades to justify premium biodiversity unit pricing
Restoration of degraded sites:
- Rebuild functional microbial communities on contaminated or compacted soils
- Establish positive plant-microbe feedbacks that self-sustain over time
- Create reference conditions for future biodiversity assessments
Climate-Driven Microbial Evolution and Survey Considerations
Drought Selection for Antibiotic Resistance
A March 2026 study published in Nature Microbiology by Caltech researchers revealed an unexpected consequence of drought: drier soils harbor significantly higher abundances of antibiotic-producing and antibiotic-resistant microbes.[3] This occurs because reduced soil moisture decreases living space, intensifying microbial competition and selecting for chemical warfare capabilities.
Geographic analysis found a strong positive correlation between regional soil aridity levels and antibiotic-resistant infection rates in hospitals, demonstrating direct climate-to-human-health connections through soil microbe evolution.[3]
For ecologists conducting Soil Microbe Memory in Drought-Resilient Biodiversity Surveys: Protocols for Ecologists Facing 2026 Climate Extremes, this finding necessitates:
⚠️ Safety protocols: Use appropriate PPE when handling soil samples from drought-prone areas
⚠️ Functional screening: Include antibiotic resistance gene profiling in eDNA analysis
⚠️ Risk assessment: Document potential human health considerations in biodiversity reports
⚠️ Management implications: Consider antibiotic resistance proliferation when designing irrigation regimes for restored habitats
Functional Shifts Parallel Community Changes
Soil microbial community structure changes occur in concert with ecosystem functioning, with direct consequences for soil carbon cycling and other biogeochemical processes.[5] This parallel shift means that compositional changes detected through eDNA surveys directly translate to functional capacity changes relevant for ecosystem service provision.
Key functional indicators to monitor:
| Functional Category | Indicator Genes/Taxa | Ecosystem Service Link |
|---|---|---|
| Carbon cycling | Cellulase, laccase genes; saprophytic fungi | Decomposition, nutrient release |
| Nitrogen cycling | nifH, amoA, nirK genes; nitrogen-fixing bacteria | Soil fertility, plant productivity |
| Stress tolerance | Heat shock proteins, osmolyte synthesis | Drought resilience, stability |
| Plant growth promotion | ACC deaminase, siderophore production | Vegetation establishment, recovery |
Regional Reference Databases for 2026 Conditions
As climate patterns shift, historical reference conditions become less reliable. Ecologists need contemporary reference databases that reflect 2026 climate realities rather than pre-industrial or even late-20th-century baselines.
Building regional reference databases:
- Identify climate-analog sites: Find locations currently experiencing conditions projected for your study area in 2030-2050
- Characterize microbial communities: Conduct comprehensive eDNA surveys of these analog sites
- Document performance metrics: Measure actual resilience (recovery rates, productivity, stability) under real-world conditions
- Establish regional standards: Develop region-specific MRI thresholds and target community compositions
- Update regularly: Reassess reference conditions every 3-5 years as climate continues changing
This approach ensures that biodiversity net gain targets reflect achievable, climate-appropriate goals rather than nostalgic recreations of past ecosystems that may no longer be viable.
Implementation Roadmap for Ecologists
Phase 1: Capacity Building (Months 1-3)
Training and equipment acquisition:
- Attend workshops on eDNA sampling techniques and bioinformatic analysis
- Establish partnerships with molecular ecology laboratories
- Invest in field sampling equipment (sterile corers, portable freezers, GPS units)
- Develop internal protocols and quality assurance procedures
Pilot project selection:
- Choose 2-3 representative sites spanning different habitat types and drought exposure histories
- Conduct parallel traditional and microbial surveys to establish baseline correlations
- Validate MRI scoring against known site resilience (historical recovery data)
Phase 2: Integration into Standard Practice (Months 4-12)
Protocol standardization:
- Incorporate eDNA sampling into all baseline biodiversity surveys
- Develop client-facing materials explaining microbial memory concepts
- Create standardized reporting templates that integrate MRI scores
- Train field staff on proper sampling techniques and chain-of-custody procedures
Data management infrastructure:
- Establish secure databases for storing eDNA sequence data
- Develop GIS layers showing regional microbial resilience patterns
- Create visualization tools for presenting microbial data to non-specialist audiences
- Implement quality control checks for laboratory results
Phase 3: Advanced Applications (Year 2+)
Predictive modeling:
- Develop site-specific resilience models incorporating microbial, plant, and climate variables
- Use machine learning to predict recovery trajectories based on baseline microbial communities
- Create decision-support tools for comparing restoration intervention options
Market development:
- Offer microbial resilience assessments as premium service for biodiversity unit sellers
- Develop certification programs for climate-adapted habitat creation
- Establish partnerships with SynCom suppliers and soil health companies
- Contribute to policy development around microbial considerations in biodiversity regulation
Case Study: Applying Microbial Memory Protocols to a Mixed Grassland Restoration
Project context: A 50-hectare former agricultural site in southeast England designated for conversion to species-rich grassland to generate biodiversity units for off-site biodiversity net gain delivery.
Baseline assessment findings:
- Traditional survey: Poor condition grassland (3 distinctiveness, 1 condition = 3 units/ha)
- Soil eDNA analysis: Low microbial diversity, dominance of agricultural-adapted taxa, MRI score = 3/10
- Drought history: Three significant droughts in past decade, slow vegetation recovery observed
Intervention strategy:
- Plant diverse seed mix (45 native species) including deep-rooted perennials
- Apply SynCom inoculant containing 15 drought-resilient rhizosphere species
- Implement adaptive grazing regime to promote root turnover and exudate production
- Establish 5-year monitoring program with annual eDNA sampling
Projected outcomes:
- Year 5 target: Moderate condition grassland (3 distinctiveness, 2 condition = 6 units/ha)
- Microbial resilience enhancement: MRI score increase to 7/10
- Climate adaptation premium: 15% uplift in unit value due to documented resilience
- Total unit generation: 300 units with enhanced marketability to climate-conscious buyers
This approach demonstrates how Soil Microbe Memory in Drought-Resilient Biodiversity Surveys: Protocols for Ecologists Facing 2026 Climate Extremes translates into tangible project value and measurable ecological outcomes.
Conclusion
The discovery that soil microbes possess ecological memory of past droughts fundamentally changes how ecologists should approach biodiversity surveys in 2026 and beyond. By integrating eDNA protocols, Microbial Resilience Index scoring, and synthetic community applications into standard practice, ecologists can quantify and enhance the climate resilience of habitats in ways that traditional surveys miss entirely.
Key action steps for immediate implementation:
✅ Begin collecting soil samples during your next biodiversity survey—even if full eDNA analysis isn't immediately available, archived samples can be analyzed later as capacity develops
✅ Document site drought history systematically, creating a database that correlates microbial community characteristics with known resilience performance
✅ Pilot SynCom applications on a subset of restoration projects to build evidence for effectiveness in your regional context
✅ Engage with policy development by contributing microbial considerations to evolving biodiversity net gain guidance and standards
✅ Build partnerships with molecular ecology labs, soil health companies, and climate scientists to access expertise and share knowledge
As climate extremes intensify throughout 2026 and beyond, the invisible world of soil microbes will increasingly determine which habitats thrive and which fail. Ecologists who embrace Soil Microbe Memory in Drought-Resilient Biodiversity Surveys: Protocols for Ecologists Facing 2026 Climate Extremes position themselves at the forefront of climate-adapted conservation practice, delivering habitats that don't just meet today's standards but remain resilient through tomorrow's challenges.
The future of biodiversity conservation lives underground—it's time our survey protocols reflected that reality.
References
[1] EGU26-22053 – https://meetingorganizer.copernicus.org/EGU26/EGU26-22053.html
[2] Managing Microbes for More Resilient Plants – https://eesa.lbl.gov/2026/02/13/managing-microbes-for-more-resilient-plants/
[3] Drought Spurs Antibiotic-Resistant Soil – https://phys.org/news/2026-03-drought-spurs-antibiotic-resistant-soil.html
[4] Global Change Biology – https://onlinelibrary.wiley.com/doi/10.1111/gcb.70441
[5] Soil Microbes Show Ecological Memory of Drought – https://communities.springernature.com/posts/soil-microbes-show-ecological-memory-of-drought


