{"cover":"Professional landscape format (1536×1024) hero image with bold text overlay 'Genetic Adaptation Tracking in Biodiversity Surveys: eDNA Protocols for Climate-Stressed Populations in 2026' in extra large 72pt white sans-serif font with dark gradient shadow, centered in upper third. Background shows split scene: left side features scientist in field gear collecting water sample from mountain stream with pipette and sample vials, right side displays DNA helix visualization with climate data overlays including temperature graphs and species distribution heat maps. Foreground includes subtle molecular structure patterns. Color palette: deep forest green, scientific blue, white text with amber accent highlights. High contrast, National Geographic editorial quality, conservation science aesthetic with modern technology integration","content":["Detailed landscape format (1536×1024) image showing close-up laboratory scene of eDNA sample processing workflow. Central focus on gloved hands pipetting water sample into PCR tubes arranged in rack, with thermal cycler machine visible in background displaying temperature readout. Left side shows filtered water samples in collection bottles with habitat labels, right side features computer monitor displaying DNA sequence chromatogram peaks and species identification results. Include magnified inset circle showing microscopic view of environmental DNA fragments. Clean scientific aesthetic with white lab surfaces, blue nitrile gloves, professional lighting. Color scheme: clinical white, laboratory blue, DNA sequence rainbow colors. Technical precision, research publication quality","Detailed landscape format (1536×1024) conceptual illustration showing climate change impact on genetic adaptation in wildlife populations. Split vertical composition: left panel depicts healthy diverse ecosystem with multiple species silhouettes (birds, amphibians, fish, mammals) connected by glowing genetic marker lines showing gene flow patterns. Right panel shows same landscape under climate stress with elevated temperature indicators, reduced species diversity, and highlighted adaptive genetic variants in surviving populations shown as bright nodes. Include DNA double helix bridge connecting both panels with mutation markers. Background transitions from lush green to drought-stressed brown landscape. Infographic style with scientific accuracy, conservation biology focus, data visualization elements including adaptation rate arrows and genetic diversity metrics","Detailed landscape format (1536×1024) field application scene showing integrated biodiversity monitoring technology in remote mountain habitat. Wide angle view of climate-stressed alpine ecosystem with visible environmental changes (receding snowline, altered vegetation zones). Foreground shows field researcher using portable TinyML device for real-time eDNA analysis at stream sampling point, with tablet displaying species detection dashboard and GPS mapping interface. Middle ground features sampling grid markers across landscape. Background shows dramatic mountain vista with weather monitoring station. Include overlay graphics showing detected species icons, genetic diversity heat map, and conservation status indicators. Color palette: natural earth tones, technology blue interface elements, amber alert highlights. Documentary photography style merged with scientific data visualization, conservation technology focus"]"}

Climate change is rewriting the genetic code of life on Earth faster than scientists can document it. A recent 56-day environmental DNA survey across western China detected nearly 400 vertebrate species spanning more than 30,000 km²—from Himalayan foothills to tropical forests—demonstrating that genetic adaptation tracking has entered a new era of precision and scale [2]. As temperatures rise and habitats shift, populations are either adapting at the genetic level or facing extinction, making Genetic Adaptation Tracking in Biodiversity Surveys: eDNA Protocols for Climate-Stressed Populations in 2026 not just a scientific advancement, but an urgent conservation necessity.
Environmental DNA (eDNA) technology has revolutionized how ecologists monitor biodiversity, offering a non-invasive window into the genetic changes occurring within climate-stressed populations. Unlike traditional survey methods that require direct observation or capture, eDNA protocols analyze genetic material shed by organisms into their environment—water, soil, or air—providing comprehensive species detection and genetic variation data simultaneously.
For developers, planners, and conservation professionals working within frameworks like Biodiversity Net Gain (BNG), understanding these advanced genetic monitoring techniques is essential. Climate-adapted populations represent irreplaceable genetic diversity that must be identified, protected, and incorporated into resilient conservation strategies.
Key Takeaways
- 🧬 eDNA surveys can detect hundreds of species across vast landscapes in weeks, identifying climate-vulnerable populations through genetic markers
- 📊 New statistical frameworks account for observation errors and model community composition changes across environmental gradients with unprecedented accuracy
- 🌡️ Genetic adaptation tracking reveals which populations possess climate-resilient traits, informing targeted conservation and habitat restoration priorities
- 💻 TinyML and optical AI technologies enable real-time genetic monitoring in remote locations without internet connectivity
- 🎯 Integration with BNG strategies ensures development projects protect genetically diverse, climate-adapted populations for long-term ecological resilience
Understanding eDNA Technology for Genetic Adaptation Tracking in Biodiversity Surveys
Environmental DNA represents genetic material released by organisms through skin cells, scales, feces, mucus, gametes, and decomposing tissue. Every species leaves a unique genetic signature in its environment, creating an invisible library of biodiversity information waiting to be decoded.
How eDNA Protocols Work in 2026
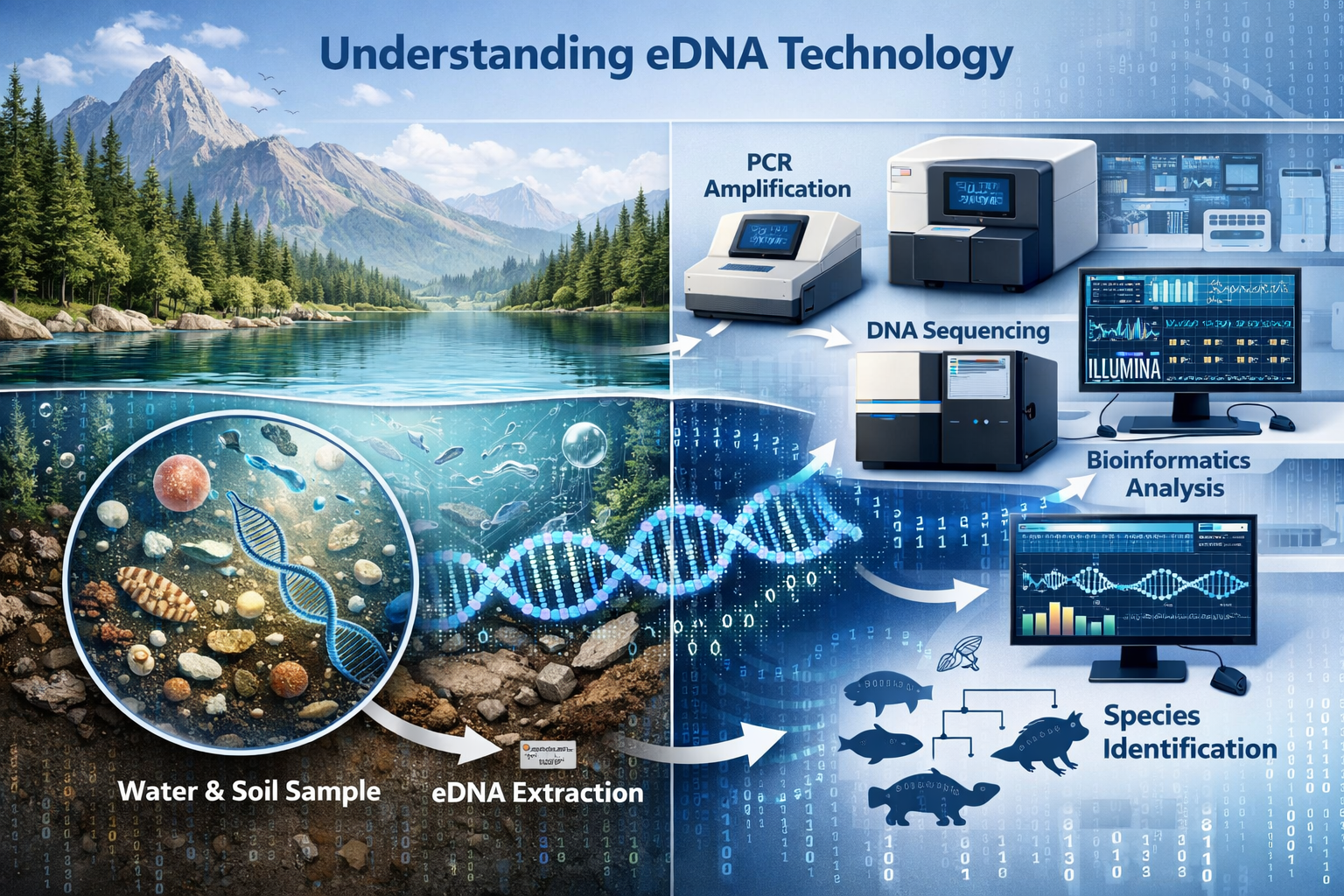
Modern eDNA protocols follow a systematic workflow:
- Sample Collection: Water, soil, or air samples are collected from target habitats using standardized methods
- DNA Extraction: Genetic material is isolated from environmental samples using specialized filtration and extraction kits
- DNA Amplification: Polymerase chain reaction (PCR) or metabarcoding techniques amplify specific genetic markers
- Sequencing: High-throughput sequencing identifies species and genetic variants present in samples
- Bioinformatic Analysis: Advanced algorithms match sequences to reference databases and identify adaptation signatures
The breakthrough in 2026 lies in the integration of genetic adaptation markers into standard eDNA protocols. Researchers can now simultaneously detect species presence and identify specific genetic variants associated with climate tolerance, disease resistance, and physiological adaptation [1].
Advanced Statistical Models for Climate-Stressed Population Assessment
Traditional eDNA studies focused primarily on species detection. However, new statistical frameworks specifically designed for eDNA metabarcoding explicitly account for observation error at multiple stages—both field sampling and laboratory procedures—and can model community composition changes across environmental gradients [2].
These models enable conservationists to:
- Predict species distribution shifts under various climate scenarios
- Identify populations with adaptive genetic variants that may serve as source populations for restoration
- Quantify genetic diversity loss in real-time across landscapes
- Validate conservation outcomes by measuring protected area effectiveness
Research has confirmed that protected areas show more occurrences of sensitive and threatened species inside boundaries and more invasive species outside—providing data-driven validation that genetic monitoring can measure real conservation impacts [2].
Climate Change Impacts on Genetic Diversity: Why Tracking Adaptation Matters
Climate change acts as a powerful selective force, favoring individuals with genetic variants that confer tolerance to heat stress, drought, altered precipitation patterns, and shifting seasonal timing. Understanding these genetic adaptations is crucial for conducting biodiversity impact assessments that account for future climate scenarios.
Identifying Climate-Vulnerable Populations Through Genomic Analysis
Recent genomic research on Populus lasiocarpa, an alpine forest species in a global biodiversity hotspot, identified western populations (primarily in the Hengduan Mountains) as most vulnerable to climate change when genetic adaptation, migration capacity, and genetic load factors are incorporated into risk predictions [1].
Key genetic indicators of climate vulnerability include:
| Genetic Factor | Climate Vulnerability Signal | Conservation Implication |
|---|---|---|
| Low genetic diversity | Limited adaptive potential | Priority for genetic rescue |
| High genetic load | Accumulation of deleterious mutations | Reduced fitness under stress |
| Adaptive haplotype blocks | Inversion polymorphisms linked to climate tolerance | Source populations for restoration |
| Restricted gene flow | Geographic isolation | Limited capacity for assisted migration |
| Local adaptation signatures | Environment-specific alleles | Populations adapted to future conditions |
Adaptive Haplotype Blocks and Climate Tolerance
Research demonstrates that haplotype blocks caused by inversion polymorphisms—which suppress recombination—are linked to enriched combinations of locally adaptive environmental variations [1]. These genetic structures act as "supergenes" that maintain beneficial combinations of traits across generations.
For practitioners implementing Biodiversity Net Gain strategies, identifying populations carrying these adaptive haplotype blocks is essential. These populations represent genetic reservoirs that may enable species persistence under future climate conditions.
"Incorporating genetic adaptation, migration, and genetic load into climate vulnerability assessments reveals hidden risks that traditional ecological models miss entirely." [1]
Implementing eDNA Protocols for Climate-Stressed Populations in 2026
The practical application of Genetic Adaptation Tracking in Biodiversity Surveys: eDNA Protocols for Climate-Stressed Populations in 2026 requires careful planning, standardized methods, and integration with existing conservation frameworks.
Four-Phase Conservation Framework with Genetic Assessment
A structured conservation protocol emphasizes four key phases [4]:
-
Documentation Phase
- Species discovery through eDNA surveys
- Specimen collection and genetic reference library development
- Baseline genetic diversity assessment
-
Assessment Phase
- Species relationships and phylogenetic analysis
- Genetic diversity quantification with climate vulnerability analysis
- Identification of adaptive genetic variants
- Risk modeling under climate scenarios
-
Monitoring Phase
- Repeated eDNA sampling across temporal scales
- Population genetic change tracking
- Habitat condition assessment
- Early warning system for genetic erosion
-
Action Phase
- Targeted habitat protection for genetically diverse populations
- Genetic rescue interventions where appropriate
- Adaptive management based on monitoring data
- Integration with BNG assessment protocols
Emerging Technologies: TinyML and Optical AI for Remote Monitoring
The 2026 Global Horizon Scan, published in Trends in Ecology & Evolution, identifies low-power Tiny Machine Learning (TinyML) devices and energy-efficient optical AI chips as imminent tools for real-time genetic and species biodiversity detection in remote, climate-stressed landscapes [3].
Advantages of TinyML for eDNA monitoring:
- ✅ No internet connection required for operation
- ✅ Ultra-low power consumption enabling solar-powered deployment
- ✅ Real-time species detection in the field
- ✅ Accessible to communities with limited digital infrastructure
- ✅ Cost-effective scaling across large landscapes
These technologies democratize genetic monitoring, enabling conservation organizations and developers planning projects to conduct comprehensive biodiversity assessments in previously inaccessible locations.
Standard Operating Procedures for eDNA Collection in Climate-Stressed Habitats
Water Sample Collection Protocol:
- Site Selection: Target areas showing climate stress indicators (temperature anomalies, altered hydrology, vegetation shifts)
- Sampling Timing: Collect during peak activity periods for target taxa
- Replication: Minimum 3 technical replicates per site
- Volume: 1-2 liters per sample for comprehensive species detection
- Filtration: 0.45-1.5 μm pore size filters to capture cellular and extracellular DNA
- Preservation: Immediate preservation in ethanol or specialized buffers
- Storage: -20°C or -80°C until extraction
Quality Control Measures:
- Field blanks to detect contamination
- Positive controls with known DNA concentrations
- Negative extraction controls
- PCR inhibition testing
- Sequencing depth standardization
Integrating Genetic Data with Biodiversity Net Gain Planning
For developers and planners working within BNG frameworks, genetic adaptation data provides critical information for creating biodiversity plans that ensure long-term ecological resilience.
Key integration points:
🎯 Baseline Assessment: Include eDNA surveys in initial site assessments to identify climate-adapted populations
🎯 Impact Prediction: Model genetic diversity loss alongside habitat loss in impact calculations
🎯 Mitigation Hierarchy: Prioritize avoidance of sites containing genetically unique or climate-adapted populations
🎯 Habitat Creation: Source propagules from genetically diverse, climate-adapted populations for restoration
🎯 Monitoring: Implement eDNA-based genetic monitoring as part of 30-year BNG management plans

Case Studies: eDNA Genetic Adaptation Tracking in Action
Western China Biodiversity Hotspot Survey
The 56-day eDNA survey across western China's biodiverse terrain demonstrates the scalability of genetic monitoring [2]. Researchers collected aquatic samples from 101 locations, detecting nearly 400 vertebrate species including mammals, birds, reptiles, amphibians, and fishes.
Key outcomes:
- Rapid assessment of biodiversity across climate gradients from Himalayan foothills to tropical forests
- Identification of species distribution patterns correlated with climate variables
- Detection of rare and cryptic species missed by traditional surveys
- Baseline genetic data for long-term climate adaptation monitoring
Protected Area Effectiveness Validation
eDNA monitoring has independently confirmed the protected area effect, showing measurably different community compositions inside versus outside conservation boundaries [2]. Protected areas demonstrated:
- Higher occurrence rates of sensitive and threatened species
- Greater genetic diversity within populations
- Lower presence of invasive species
- Stronger signatures of climate-adapted genotypes
This validation demonstrates that eDNA protocols can provide objective, quantifiable metrics for measuring conservation outcomes required in BNG frameworks.
Alpine Forest Climate Vulnerability Assessment
Genomic analysis of Populus lasiocarpa populations revealed that western populations face highest climate vulnerability when genetic factors are considered [1]. Traditional ecological niche models would have missed this vulnerability, potentially leading to inadequate conservation prioritization.
The study identified:
- Specific adaptive haplotype blocks associated with temperature tolerance
- Geographic patterns of genetic load accumulation
- Migration capacity limitations under rapid climate change
- Priority populations for ex-situ conservation and genetic rescue
Practical Applications for Developers and Conservation Professionals
Pre-Development eDNA Assessments
Before initiating development projects, conducting eDNA surveys provides comprehensive baseline data that satisfies regulatory requirements while identifying conservation priorities:
Benefits for developers:
- Faster, more cost-effective than traditional survey methods
- Year-round sampling capability (not constrained by species activity periods)
- Detection of protected species that might otherwise be missed
- Defensible data for planning applications
- Reduced project delays from unexpected species discoveries
Integration with BNG requirements:
Genetic diversity metrics can be incorporated into biodiversity unit calculations, potentially increasing the value of habitats containing climate-adapted populations and incentivizing their protection.
Habitat Banking and Genetic Diversity Premiums
Landowners creating habitat banks can leverage genetic adaptation data to demonstrate higher ecological value. Sites containing populations with documented climate-adapted genotypes may command premium prices in biodiversity credit markets.
Value-adding factors:
- Documented genetic diversity above regional averages
- Presence of adaptive haplotype blocks
- Source populations for climate-resilient restoration
- Connectivity to other genetically diverse populations
- Long-term climate suitability modeling
Monitoring and Adaptive Management
Long-term eDNA monitoring enables adaptive management by tracking genetic changes in response to:
- Climate change progression
- Habitat restoration effectiveness
- Management intervention outcomes
- Invasive species impacts
- Disease emergence
Regular eDNA sampling (quarterly or annually) provides early warning signals of genetic erosion, enabling proactive conservation responses before population viability is compromised.
Future Directions: Emerging Technologies and Methodologies
Portable Sequencing and Field-Based Analysis
The miniaturization of DNA sequencing technology continues to advance. Portable sequencers like Oxford Nanopore's MinION enable field-based genetic analysis, reducing the time from sample collection to species identification from weeks to hours.
2026 capabilities:
- Real-time species detection during field surveys
- Immediate identification of protected or invasive species
- On-site genetic diversity assessment
- Rapid response to biosecurity threats
- Community-based monitoring programs
Integration with Remote Sensing and AI
The convergence of eDNA data with satellite remote sensing and artificial intelligence creates powerful predictive models for biodiversity distribution and climate adaptation potential [3].
Emerging applications:
- Habitat suitability modeling incorporating genetic adaptation capacity
- Predictive mapping of climate-vulnerable populations
- Automated species detection from environmental samples
- Integration with biodiversity credit trading platforms
Standardization and Regulatory Integration
As eDNA methods mature, regulatory frameworks are beginning to accept genetic monitoring data for compliance purposes. The UK's Environment Act 2021 and associated BNG regulations provide opportunities for integrating genetic diversity metrics into statutory requirements.
Anticipated developments:
- Standardized eDNA protocols for BNG assessments
- Genetic diversity thresholds in habitat quality metrics
- Mandatory genetic monitoring for large-scale developments
- Integration with national biodiversity databases
Challenges and Limitations of eDNA Genetic Adaptation Tracking
Despite remarkable advances, several challenges remain:
Technical Limitations
- Reference database gaps: Many species lack comprehensive genetic reference data, limiting identification accuracy
- Quantification challenges: Converting eDNA sequence reads to absolute abundance estimates remains difficult
- Temporal dynamics: eDNA persistence varies by environment, complicating interpretation
- Contamination risks: Stringent protocols required to prevent false positives
Interpretation Complexities
- Presence vs. viability: eDNA confirms presence but not population health or reproductive success
- Genetic adaptation vs. plasticity: Distinguishing genetic adaptation from phenotypic plasticity requires careful analysis
- Spatial resolution: eDNA signals can travel via water flow, obscuring precise source locations
Cost and Accessibility
While costs continue to decline, comprehensive eDNA genetic adaptation tracking remains more expensive than traditional presence/absence surveys. However, when considering the breadth of data obtained and the elimination of multiple season-specific surveys, eDNA often proves cost-effective for developers planning projects.
Conclusion
Genetic Adaptation Tracking in Biodiversity Surveys: eDNA Protocols for Climate-Stressed Populations in 2026 represents a paradigm shift in conservation science and biodiversity assessment. As climate change accelerates, identifying populations with adaptive genetic variants becomes not just scientifically interesting but essential for effective conservation and development planning.
The integration of eDNA genetic monitoring with frameworks like Biodiversity Net Gain creates unprecedented opportunities to build ecological resilience into development projects. By identifying and protecting climate-adapted populations, developers and conservation professionals can ensure that biodiversity gains persist across decades of environmental change.
Actionable Next Steps
For Developers and Planners:
- Incorporate eDNA surveys into baseline biodiversity assessments for all major projects
- Request genetic diversity analysis as part of ecological impact assessments
- Prioritize avoidance of sites containing genetically unique or climate-adapted populations
- Source restoration materials from genetically diverse, climate-adapted populations
- Implement genetic monitoring as part of long-term management plans
For Conservation Professionals:
- Establish baseline genetic databases for priority species and habitats
- Develop standardized eDNA protocols specific to regional climate stressors
- Integrate genetic metrics into habitat quality assessments
- Build partnerships with genetic laboratories and bioinformatic specialists
- Advocate for regulatory recognition of genetic diversity in conservation planning
For Landowners:
- Conduct eDNA assessments to document genetic diversity on your property
- Explore premium pricing for biodiversity units from genetically diverse habitats
- Participate in genetic monitoring programs that add value to conservation land
- Consider genetic connectivity when planning habitat management
The future of biodiversity conservation lies in understanding not just which species are present, but whether those populations possess the genetic capacity to adapt to our rapidly changing world. Environmental DNA technology provides the tools to answer this critical question at scale, enabling evidence-based decisions that protect both nature and development interests.
As we navigate the challenges of climate change and biodiversity loss, genetic adaptation tracking through eDNA protocols offers a science-based pathway toward resilient ecosystems and sustainable development. The technology exists, the methods are proven, and the time to act is now.
References
[1] Molecular Biology and Evolution – https://academic.oup.com/mbe/article/42/7/msaf116/8148825
[2] Closing Gap Between Biodiversity Commitments And Measuring Nature – https://sps.columbia.edu/news/closing-gap-between-biodiversity-commitments-and-measuring-nature
[3] Whats Next For Biodiversity Conservation Insights From The 2026 Horizon Scan – https://www.unep-wcmc.org/en/news/whats-next-for-biodiversity-conservation-insights-from-the-2026-horizon-scan
[4] NCBI Biodiversity Conservation – https://pmc.ncbi.nlm.nih.gov/articles/PMC12846871/


